Differences in cortical and white matter myelination: A challenge for MRI myelin biomarkers
Myelin, the fatty axon-insulating substance in the brain, is composed of a large variety of lipids, proteins and trapped water. It is the main source of contrast in magnetic resonance images (MRI) of the human brain. All MRI parameters, including longitudinal and transverse relaxation rates (R1, R2, R2*), proton density (PD), magnetisation transfer (MT) and magnetic susceptibility are sensitive to tissue myelination due to different biophysical mechanisms. Therefore, quantitative MR parameters can provide unique in vivo information on brain development, cortical myeloarchitecture, plasticity and neurodegeneration.
The mechanisms underlying the sensitivity of MRI parameters to tissue myelination are only partly understood and the validation of MRI-based myelin biomarkers is difficult due to the lack of methods for histological myelin quantification. Classical histological methods (stainings and immunohistochemistry) do not provide quantitative information on myelin content and reflect only some of the various myelin components. Recently developed advanced methods for lipid quantification promise to overcome this limitation. In this project, we explore lipid imaging using matrix-assisted laser desorption/ionization (MALDI) as a method for validating MRI-based myelin biomarkers, and we compare it to multiple histological myelin stains in post-mortem human brain tissue samples. The following two figures show some results we have achieved so far.
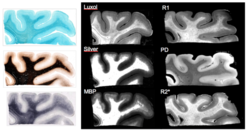
Figure 1 illustrates that all myelin stains and all quantitative MR parameters we investigated provide high contrast between white and grey matter, with white matter having higher myelin content. However, the patterns of myelination revealed in the cortex differed between stains and MRI parameters. A gradient of myelin content from the pial surface to white matter boundary was prominent in the Silver stain and anti-myelin-basic-protein immunohistochemistry, resembling the patterns observed in R2* and PD maps. Hyperintensity of cortical layer IV was observed in the Luxol Fast Blue staining, corresponding to the pattern found in the R1 maps. Our findings indicate that different histological stains for myelin detection provide similar, but distinct information on tissue myelination, particularly in the cortex. Differences between histological methods potentially correspond to differences in specificity between MRI-based myelin markers.
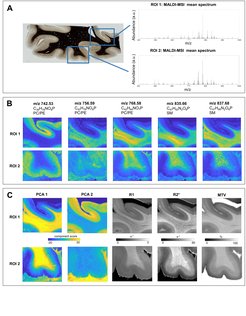
Figure 2 shows an average MALDI mass spectrum in human brain tissue with peaks corresponding to different lipid classes. Example lipid maps for selected m/z ratios are also provided. MALDI revealed distinct distributions of lipid components in white and grey matter, cortical layers and in the optic radiation. This is important for the interpretation of R1- and MT-based myelin biomarkers, since we know that lipid composition can influence relaxation properties of myelinated fibers. Combining the information from all lipid maps with a principal component analysis suggests the presence of two main lipid components (indicated as PCA1 and PCA2). Lipids of the first component are predominantly located in white matter, also demonstrating an intra-cortical gradient, and most likely reflect local myelin content. The lipids of the second components are mostly localized in the primary visual cortex. We show the component maps next to the quantitative MRI maps.
Our results so far suggest that difference in lipid composition of the cortex and in white matter pathways should be taken into account when interpreting MRI-based maps of brain myelination and that a principal component analysis of MALDI-MSI lipid maps can be used to obtain a histological myelin biomarker for the validation of quantitative MRI parameters.